Publications
Highlights
(For a full list see below or go to Google Scholar)
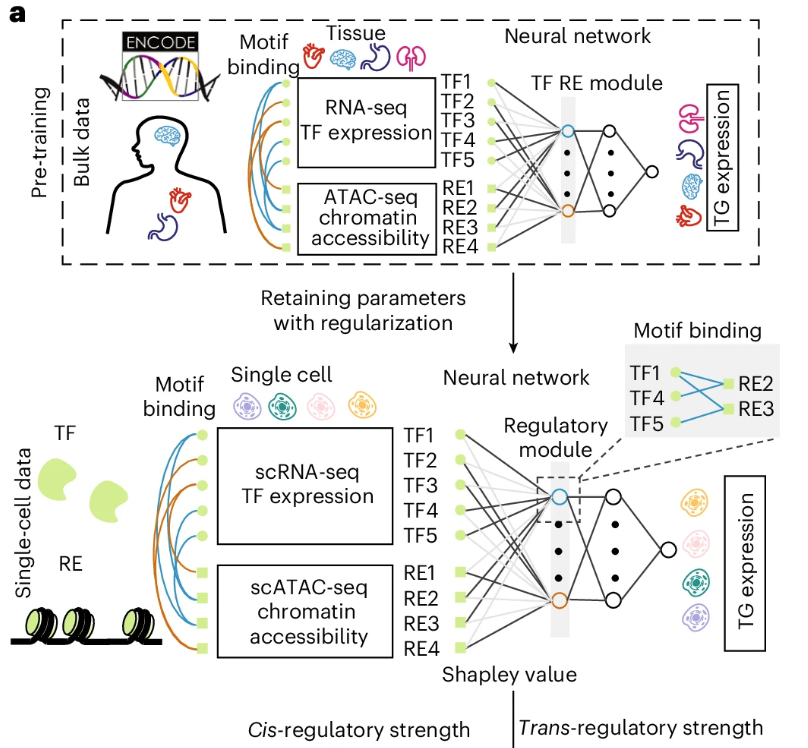
We present LINGER, a machine-learning method to infer gene regulatory networks from single-cell paired data. LINGER incorporates atlas-scale external bulk data across diverse cellular contexts and prior knowledge of transcription factor motifs as a manifold regularization to achieve high-accuracy inference.
Qiuyue Yuan and Zhana Duren
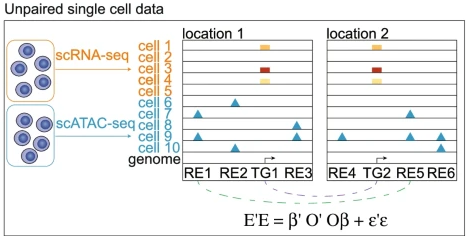
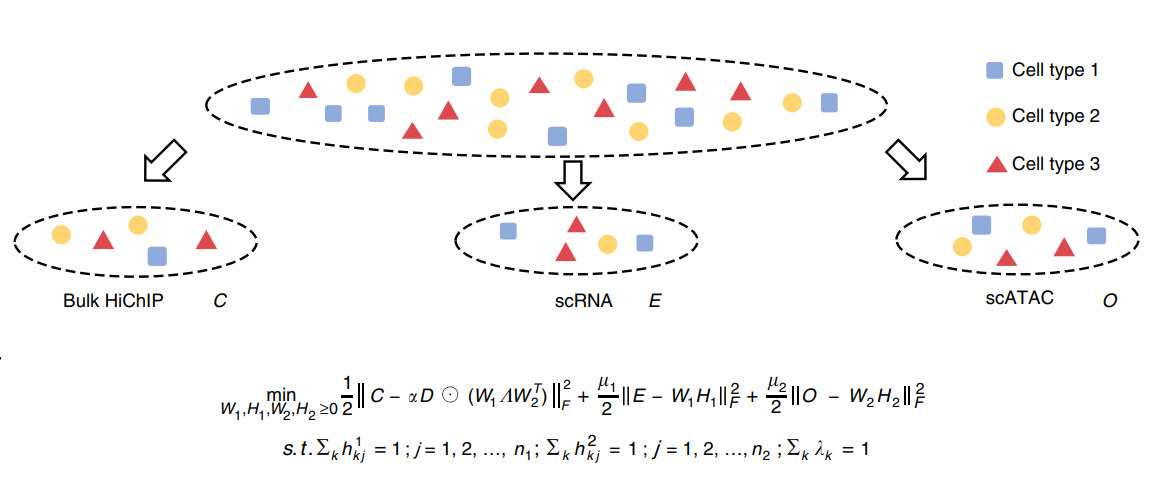
We propose UnpairReg for regression analysis on unpaired observations to integrate single-cell multi-omics data. UnpairReg provides accurate estimations of gene expression from chromatin accessibility and infers regulatory networks consistent with eQTL mapping, improving cell type identification accuracy.
Qiuyue Yuan and Zhana Duren
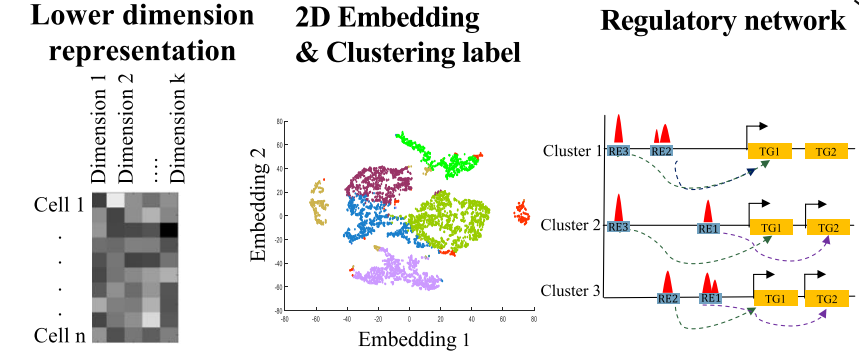
We developed scREG, a dimension reduction methodology based on the concept of cis-regulatory potential for single-cell multiome data. This framework enables the construction of subpopulation-specific regulatory networks and provides a comprehensive R package for advanced multi-omics data integration and analysis.
Zhana Duren, Fengge Chang, Fnu Naqing, Jingxue Xin, Qiao Liu, and Wing Hung Wong
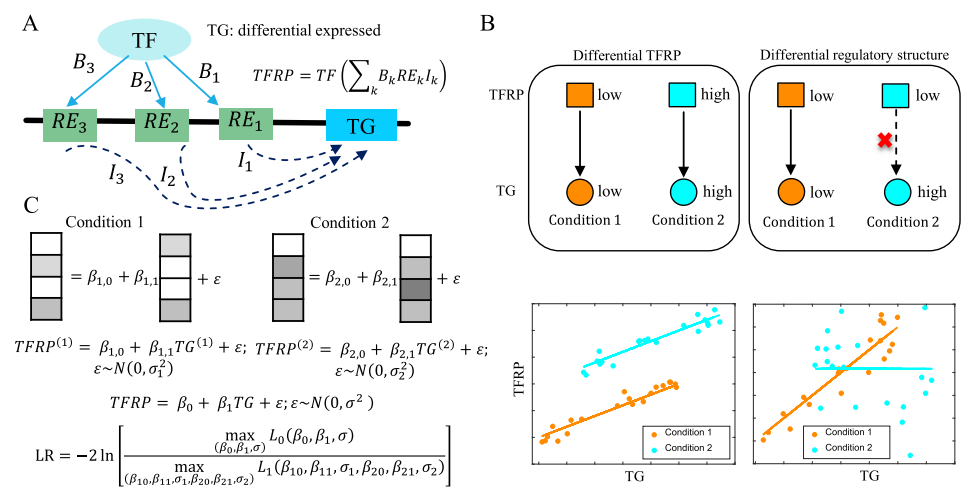
We introduce sc-compReg, a computational method designed for the comparative analysis of gene expression regulatory networks between two biological conditions. By integrating scRNA-seq and scATAC-seq data, it identifies key regulatory changes driving transitions between different cellular states or treatments.
Zhana Duren, Sophia Lu, Joseph G. Arthur, Preyas Shah, Jingxue Xin, Francesca Meschi, Miranda Lin Li, Corey M. Nemec, Yifeng Yin, and Wing Hung Wong
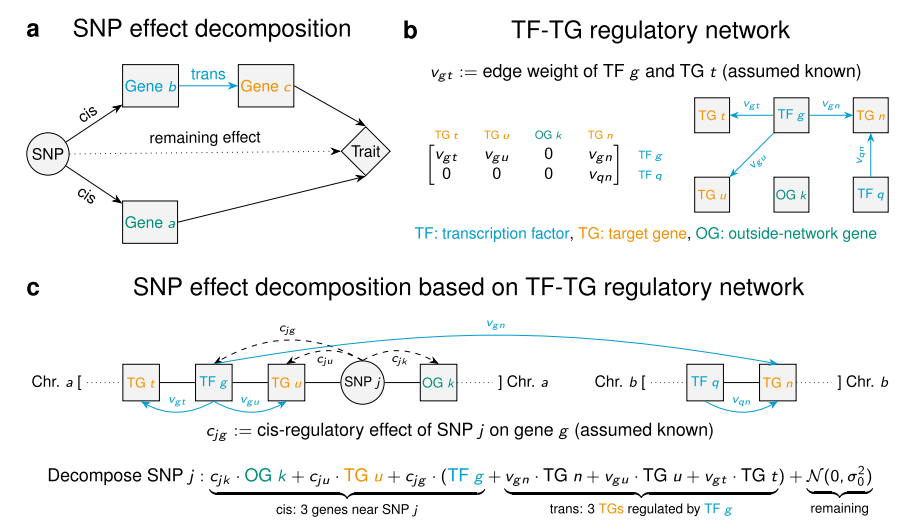
We developed a Bayesian framework that integrates GWAS summary statistics with regulatory networks to infer genetic enrichments and associations. Our method explicitly models network topology to assess enrichments and leverages this information to improve the identification of trait-associated genetic variants.
Xiang Zhu, Zhana Duren, Wing Hung Wong
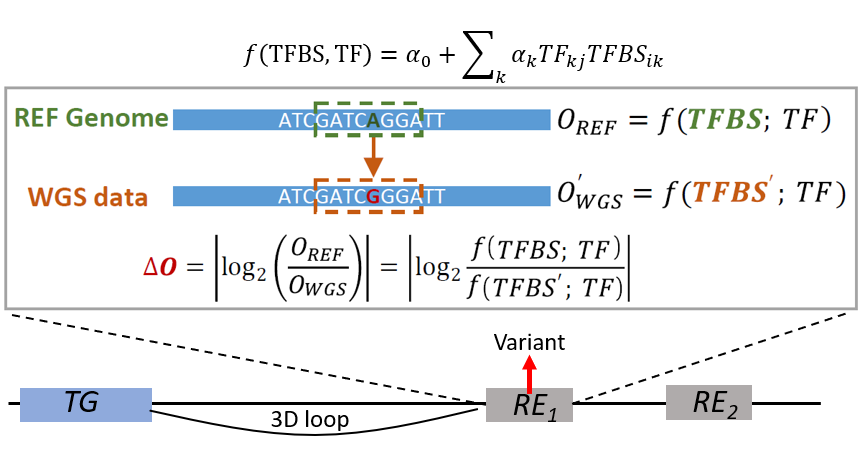
This work presents a model that learns the dependence of regulatory element accessibility on DNA sequences and TF expression. By combining sequence information with context-specific data, the approach prioritizes noncoding variants and regulatory elements, facilitating the interpretation of GWAS risk loci.
Wenran Li, Zhana Duren, Rui Jiang, Wing Hung Wong
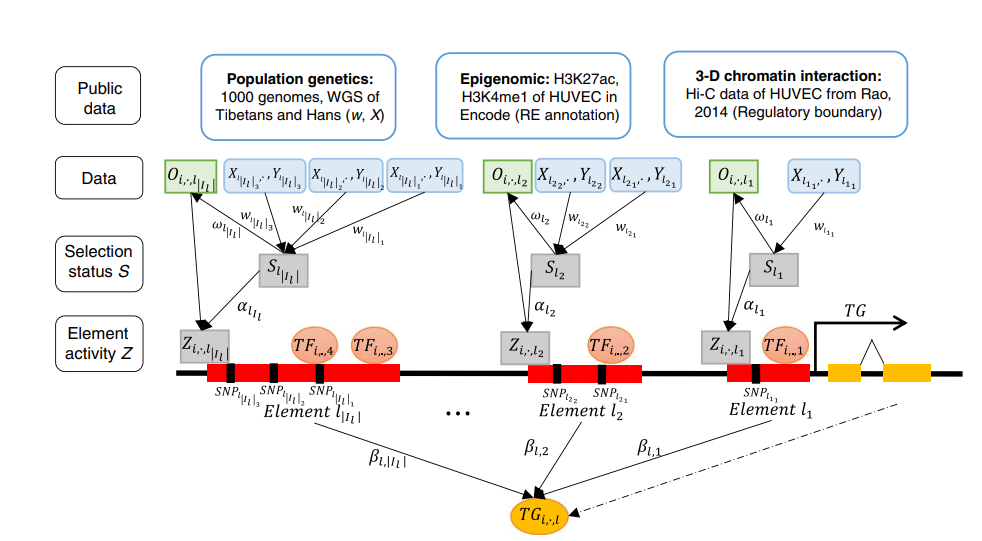
We developed vPECA to identify active selected regulatory elements and associated networks in hypoxia adaptation. We discovered three causal SNPs of EPAS1 in Tibetans that decrease chromatin accessibility and weaken TF binding strength, providing a mechanistic understanding of high-altitude adaptation.
Jingxue Xin, Hui Zhang, Yaoxi He, Zhana Duren, Caijuan Bai, Lang Chen, Xin Luo, Dong-Sheng Yan, Chaoyu Zhang, Xiang Zhu, Qiuyue Yuan, Zhanying Feng, Chaoying Cui, Xuebin Qi, Wing Hung Wong, Yong Wang, Bing Su
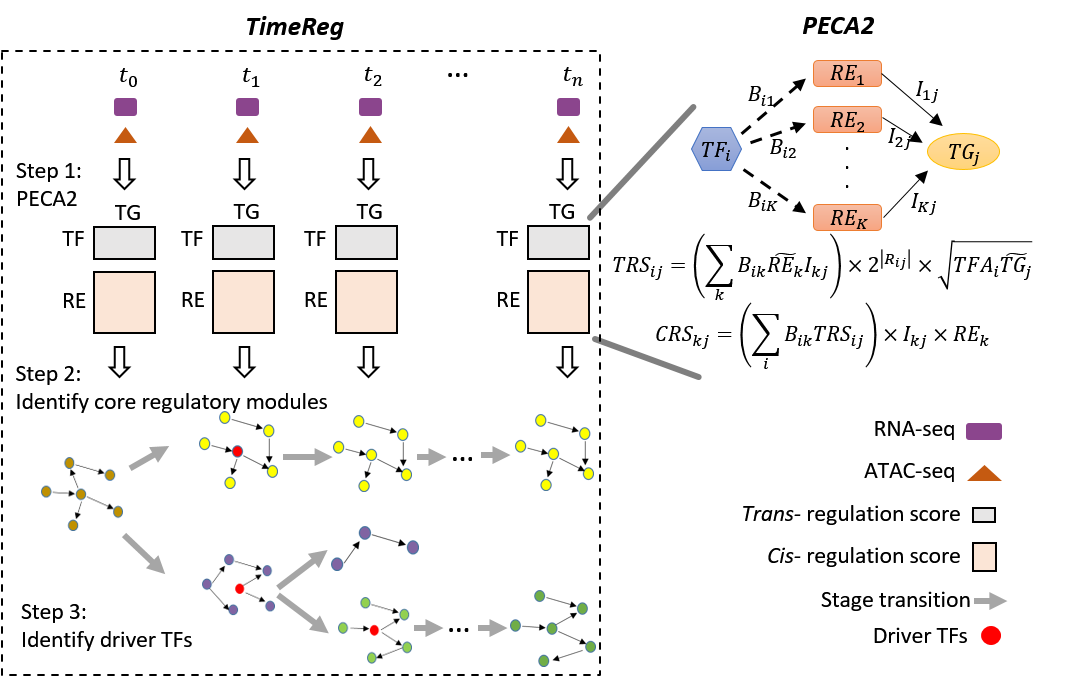
We developed TimeReg to prioritize regulatory elements and extract core regulatory modules from time-course data. TimeReg identifies key regulators driving cellular state changes and establishes causal connections between regulatory modules across different time points.
Zhana Duren, Xi Chen, Jingxue Xin, Yong Wang, Wing Hung Wong
We introduce DC3 (De-Convolution and Coupled-Clustering) for the joint analysis of bulk 3D chromatin data and single-cell genomics. DC3 simultaneously identifies distinct subpopulations, assigns single cells to clusters, and deconvolves bulk data into subpopulation-specific profiles for heterogeneous cell populations.
Wanwen Zeng, Xi Chen, Zhana Duren, Yong Wang, Rui Jiang, Wing Hung Wong
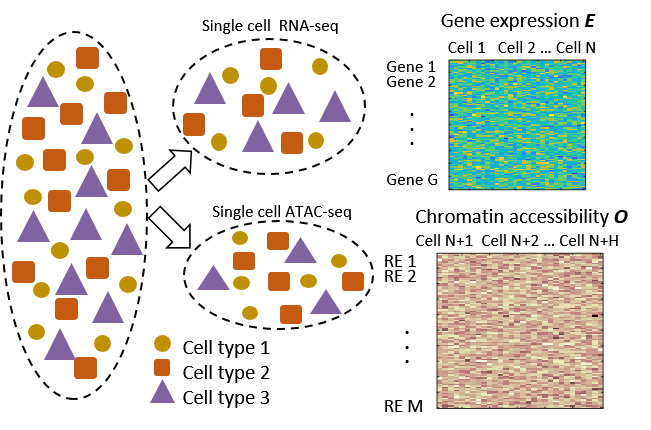
We developed a coupled nonnegative matrix factorization approach for “coupled clustering” in heterogeneous single-cell datasets. This method enables the joint analysis of different cellular features generated from different samples within the same biological mixture.
Zhana Duren, Xi Chen, Mahdi Zamanighomi, Wanwen Zeng, Ansuman T. Satpathy, Howard Y. Chang, Yong Wang, Wing Hung Wong
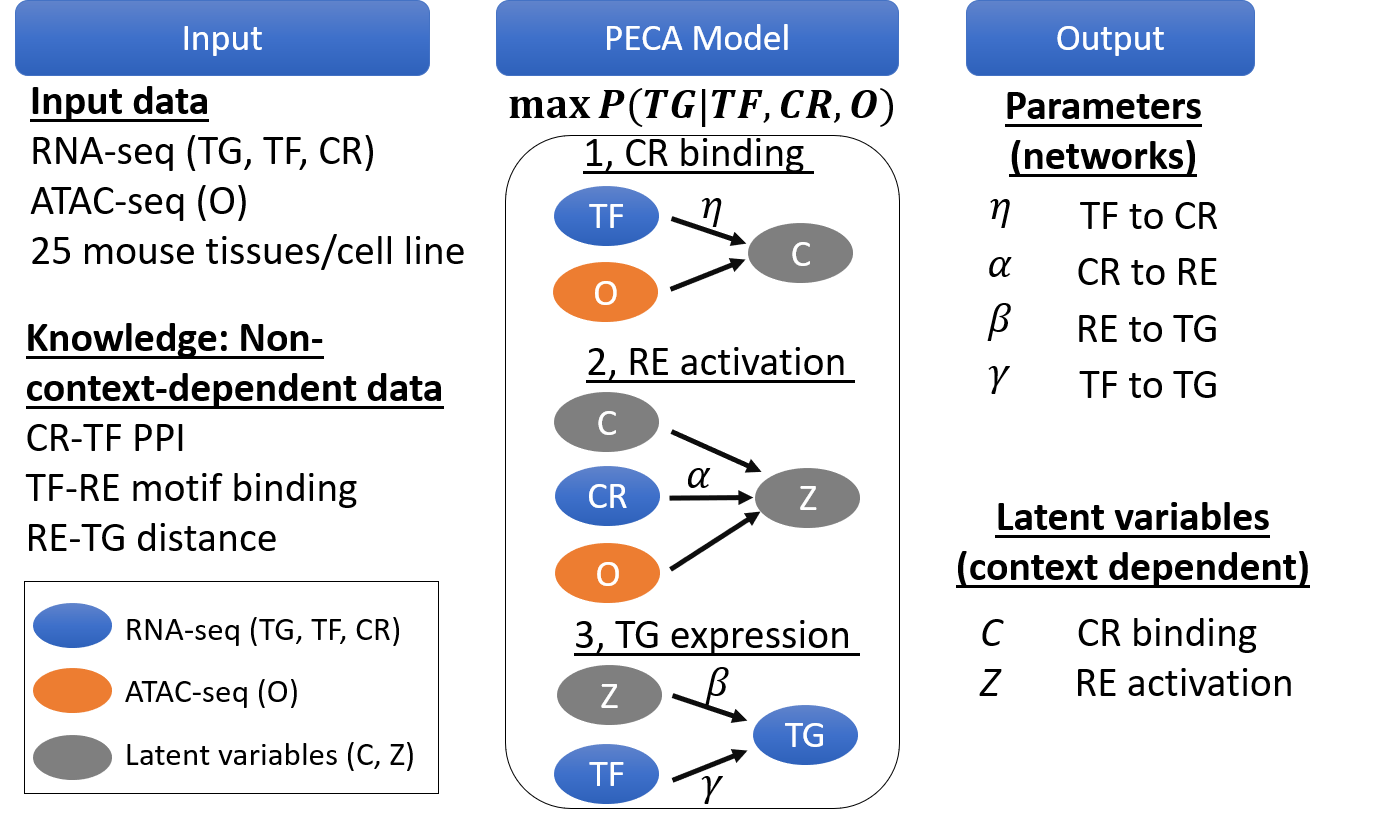
PECA is a mechanistic model that infers gene regulatory networks from paired expression and accessibility data. By modeling matched data across diverse contexts, PECA recovers missing information on binding locations and chromatin states, achieving accurate inference of gene regulatory relations and structure.
Zhana Duren, Xi Chen, Rui Jiang, Yong Wang, Wing Hung Wong
Full List
Integrated single-cell multiomic profiling of caudate nucleus suggests key mechanisms in alcohol use disorder
Nicholas C Green, Hongyu Gao, Xiaona Chu, Qiuyue Yuan, Patrick McGuire, Dongbing Lai, Guanglong Jiang, Xiaoling Xuei, Jill L Reiter, Julia Stevens, Greg T Sutherland, Alison M Goate, Zhiping P Pang, Paul A Slesinger, Ronald P Hart, Jay A Tischfield, Arpana Agrawal, Yue Wang, Zhana Duren, Howard J Edenberg, Yunlong Liu
Nature Communications 2025
A distinct population of CD8+ T cells expressing CD39 and CD73 accumulates with age and supports cancer progression
Monica Bodogai, Bongsoo Park, Fatima-Zohra Braikia, Fnu Naqing, Konda Kumaraswami, Chen Chen, Emeline Ragonnaud, Sharon Stack, Steffen Ormanns, Michael Günther, Hellen Ishikawa-Ankerhold, Supriyo De, Luigi Ferrucci, Ranjan Sen, Zhana Duren, Isabel Beerman, Arya Biragyn
Nature Aging 2025
Modeling combinatorial regulation from single-cell multi-omics provides regulatory units underpinning cell type landscape using cRegulon
Zhanying Feng, Xi Chen, Zhana Duren , Jingxue Xin, Hao Miao, Qiuyue Yuan , Yong Wang, Wing Hung Wong
Genome Biology 2025
Epiregulon: Single-cell transcription factor activity inference to predict drug response and drivers of cell states
Tomasz Włodarczyk, Aaron Lun, Diana Wu, Minyi Shi, Xiaofen Ye, Shreya Menon, Shushan Toneyan, Kerstin Seidel, Liang Wang, Jenille Tan, Shang-Yang Chen, Timothy Keyes, Aleksander Chlebowski, Adrian Waddell, Wei Zhou, Yangmeng Wang, Qiuyue Yuan, Yu Guo, Liang-Fu Chen, Bence Daniel, Antonina Hafner, Meng He, Alejandro Chibly, Yuxin Liang, Zhana Duren, Ciara Metcalfe, Marc Hafner, Christian W Siebel, M Ryan Corces, Robert Yauch, Shiqi Xie, Xiaosai Yao
Nature Communications 2025
Inferring gene regulatory networks from single-cell multiome data using atlas-scale external data
Qiuyue Yuan and Zhana Duren
Nature Biotechnology 2024
Joint inference of clonal structure using single-cell genome and transcriptome sequencing data
Xiangqi Bai, Zhana Duren, Lin Wan, Li C Xia
NAR Genomics and Bioinformatics 2024
Heritability enrichment in context-specific regulatory networks improves phenotype-relevant tissue identification
Zhanying Feng, Zhana Duren, Jingxue Xin, Qiuyue Yuan , Yaoxi He, Bing Su, Wing Hung Wong, and Yong Wang
eLife 2022
Human genetic variants associated with COVID-19 severity are enriched in immune and epithelium regulatory networks
Zhanying Feng, Xianwen Ren, Zhana Duren, and Yong Wang
Phenomics 2022
Integration of single-cell multi-omics data by regression analysis on unpaired observations
Qiuyue Yuan and Zhana Duren
Genome Biology 2022
Regulatory analysis of single cell multiome gene expression and chromatin accessibility data with scREG
Zhana Duren, Fengge Chang, Fnu Naqing, Jingxue Xin, Qiao Liu, and Wing Hung Wong
Genome Biology 2022
Sc-compReg enables the comparison of gene regulatory networks between conditions using single-cell data
Zhana Duren, Sophia Lu, Joseph G. Arthur, Preyas Shah, Jingxue Xin, Francesca Meschi, Miranda Lin Li, Corey M. Nemec, Yifeng Yin, and Wing Hung Wong
Nature communications 2021
Dynamic chromatin regulatory landscape of human CAR T cell exhaustion
David G. Gennert, Rachel C. Lynn, Jeff M. Granja, Evan W. Weber, Maxwell R. Mumbach, Yang Zhao, Zhana Duren, Elena Sotillo, William J. Greenleaf, Wing H. Wong, Ansuman T. Satpathy, Crystal L. Mackall, and Howard Y. Chang
PNAS 2021
MIMIC: an optimization method to identify cell type-specific marker panel for cell sorting
Meng Zou, Zhana Duren, Qiuyue Yuan, Henry Li, Andrew Paul Hutchins, Wing Hung Wong, Yong Wang
Briefings in Bioinformatics 2021
hReg-CNCC reconstructs a regulatory network in human cranial neural crest cells and annotates variants in a developmental context
Zhanying Feng, Zhana Duren, Ziyi Xiong, Sijia Wang, Fan Liu, Wing Hung Wong, Yong Wang
Communications biology 2021
Modeling regulatory network topology improves genome-wide analyses of complex human traits
Xiang Zhu, Zhana Duren, Wing Hung Wong
Nature communications 2021
GuidingNet: revealing transcriptional cofactor and predicting binding for DNA methyltransferase by network regularization
Lixin Ren, Caixia Gao, Zhana Duren, Yong Wang
Briefings in Bioinformatics 2021
A method for scoring the cell type-specific impacts of noncoding variants in personal genomes
Wenran Li, Zhana Duren, Rui Jiang, Wing Hung Wong
PNAS 2020
Chromatin accessibility landscape and regulatory network of high-altitude hypoxia adaptation
Jingxue Xin, Hui Zhang, Yaoxi He, Zhana Duren, Caijuan Bai, Lang Chen, Xin Luo, Dong-Sheng Yan, Chaoyu Zhang, Xiang Zhu, Qiuyue Yuan, Zhanying Feng, Chaoying Cui, Xuebin Qi, Wing Hung Wong, Yong Wang, Bing Su
Nature communications 2020
Time course regulatory analysis based on paired expression and chromatin accessibility data
Zhana Duren, Xi Chen, Jingxue Xin, Yong Wang, Wing Hung Wong
Genome Research 2020
Integrated functional genomic analyses of Klinefelter and Turner syndromes reveal global network effects of altered X chromosome dosage
Xianglong Zhang, David Hong, Shining Ma, Thomas Ward, Marcus Ho, Reenal Pattni, Zhana Duren, Atanas Stankov, Sharon Bade Shrestha, Joachim Hallmayer, Wing Hung Wong, Allan L Reiss, Alexander E Urban
PNAS 2020
Modeling regulatory network topology improves genome-wide analyses of complex human traits
Xiang Zhu, Zhana Duren, Wing Hung Wong
bioRxiv 2020
DC3 is a method for deconvolution and coupled clustering from bulk and single-cell genomics data
Wanwen Zeng, Xi Chen, Zhana Duren, Yong Wang, Rui Jiang, Wing Hung Wong
Nature Communications 2019
Hierarchical graphical model reveals HFR1 bridging circadian rhythm and flower development in Arabidopsis thaliana
Zhana Duren, Yaling Wang, Jiguang Wang, Xing-Ming Zhao, Le Lv, Xiaobo Li, Jingdong Liu, Xin-Guang Zhu, Luonan Chen, Yong Wang
NPJ systems biology and applications 2019
TFAP2C-and p63-dependent networks sequentially rearrange chromatin landscapes to drive human epidermal lineage commitment
Lingjie Li, Yong Wang, Jessica L Torkelson, Gautam Shankar, Jillian M Pattison, Hanson H Zhen, Fengqin Fang, Zhana Duren, Jingxue Xin, Sadhana Gaddam, Sandra P Melo, Samantha N Piekos, Jiang Li, Eric J Liaw, Lang Chen, Rui Li, Marius Wernig, Wing H Wong, Howard Y Chang, Anthony E Oro
Cell Stem Cell 2019
Integrative analysis of single-cell genomics data by coupled nonnegative matrix factorizations
Zhana Duren, Xi Chen, Mahdi Zamanighomi, Wanwen Zeng, Ansuman T. Satpathy, Howard Y. Chang, Yong Wang, Wing Hung Wong
PNAS 2018
Unsupervised clustering and epigenetic classification of single cells
Mahdi Zamanighomi, Zhixiang Lin, Timothy Daley, Xi Chen, Zhana Duren, Alicia Schep, William J Greenleaf, Wing Hung Wong
Nature communications 2018
Modeling gene regulation from paired expression and chromatin accessibility data
Zhana Duren, Xi Chen, Rui Jiang, Yong Wang, Wing Hung Wong
PNAS 2017
A systematic method to identify modulation of transcriptional regulation via chromatin activity reveals regulatory network during mESC differentiation
Zhana Duren, Yong Wang
Scientific reports 2016
A community effort to assess and improve drug sensitivity prediction algorithms
James C Costello, Laura M Heiser, Elisabeth Georgii et. al
Nature biotechnology 2014
Sparse hyperspectral unmixing using an approximate L0 norm
Wei Tang, Zhenwei Shi, Zhana Duren
Optik 2014
Subspace matching pursuit for sparse unmixing of hyperspectral data
Zhenwei Shi, Wei Tang, Zhana Duren, Zhiguo Jiang
IEEE Transactions on Geoscience and Remote Sensing 2013
Inferring gene regulatory network for cell reprogramming
Duren Zhana , Wang Yong, Saito Shigeru, Horimoto Katsuhisa
Control Conference (CCC), 2012 31st Chinese
Clinical data analysis reveals three subytpes of gastric cancer
Xinxin Wang, Zhana Duren, Chao Zhang, Lin Chen, Yong Wang
2012 IEEE 6th International Conference on Systems Biology (ISB)