Welcome to the Duren Lab
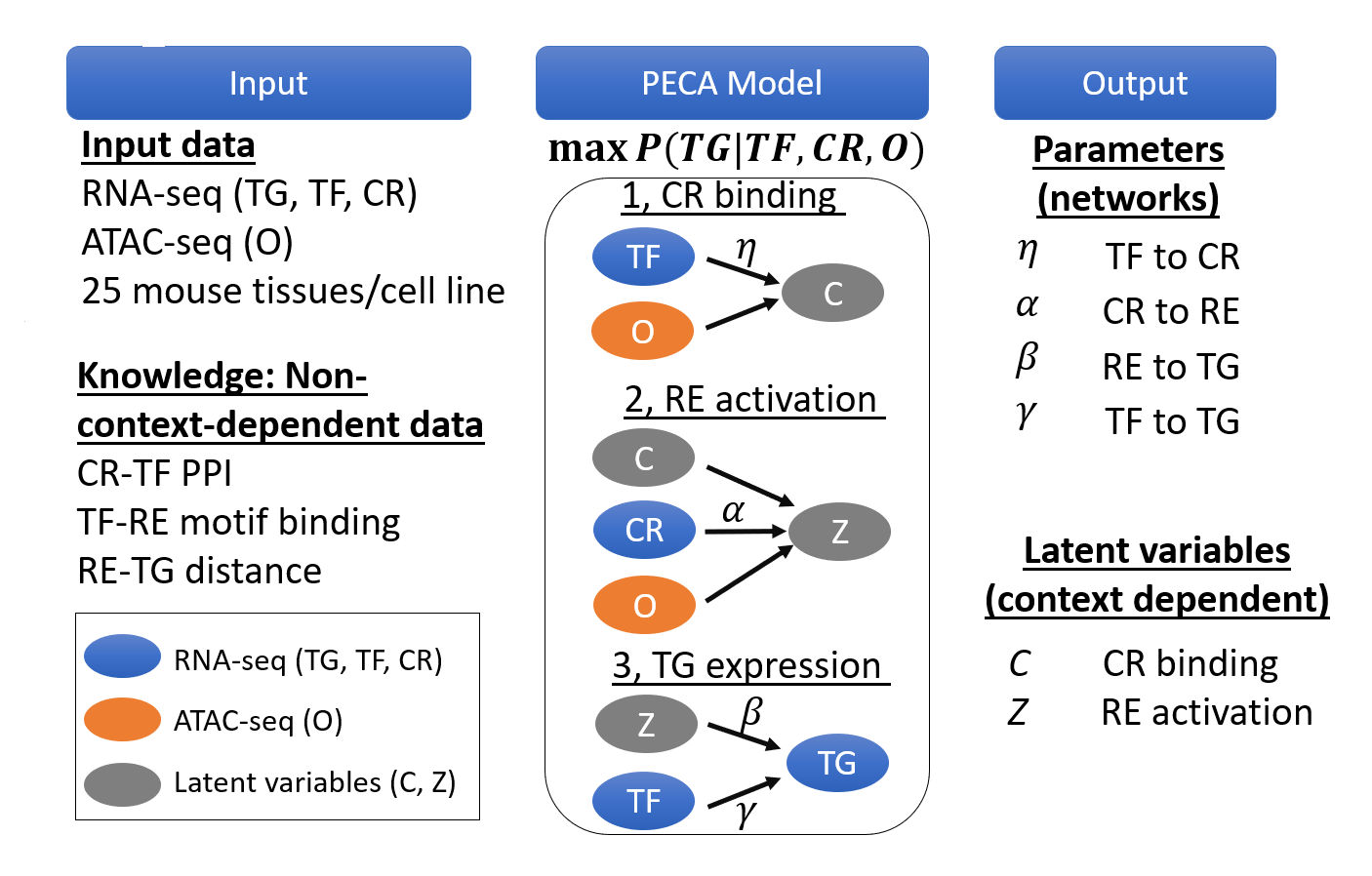
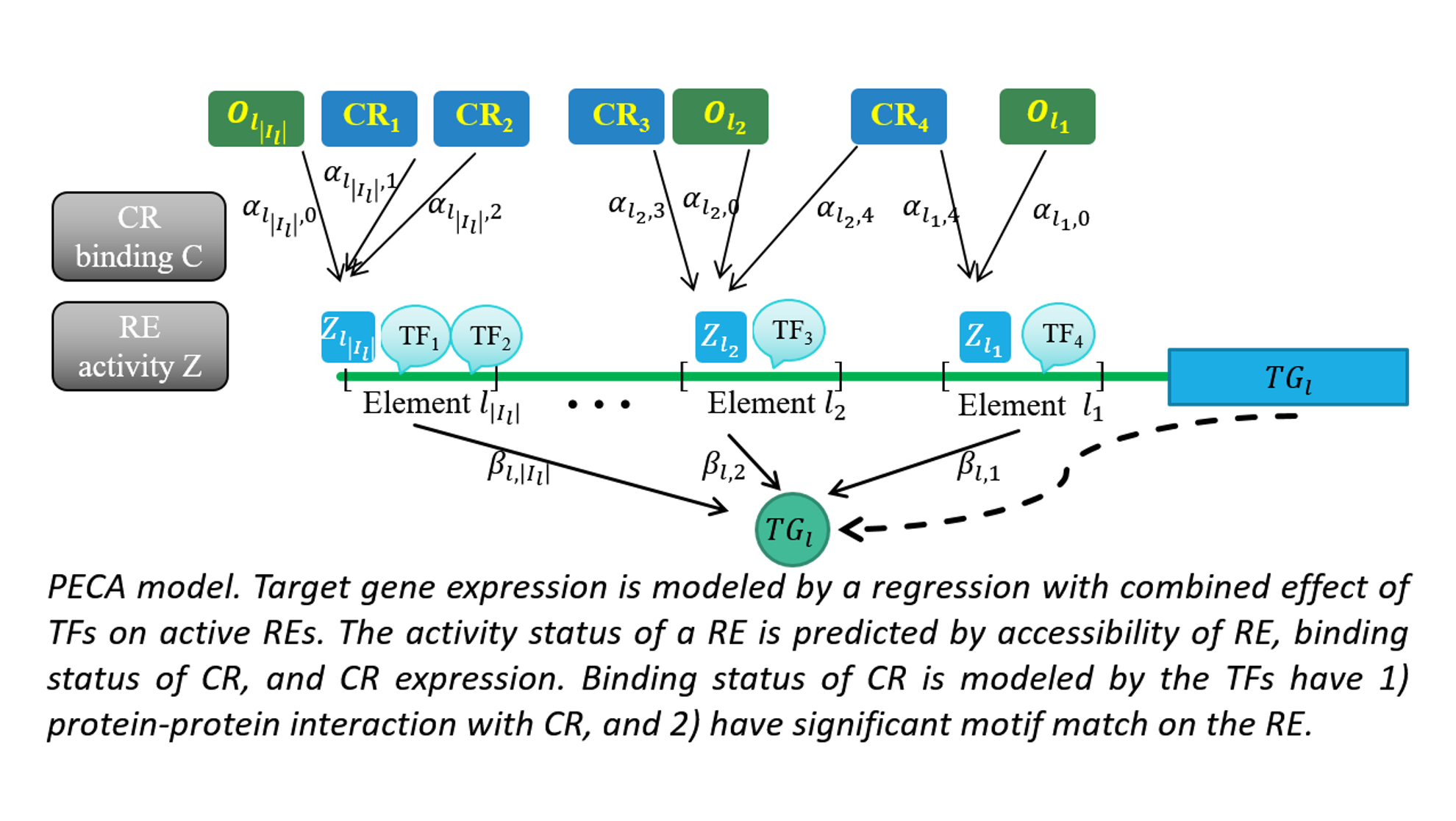
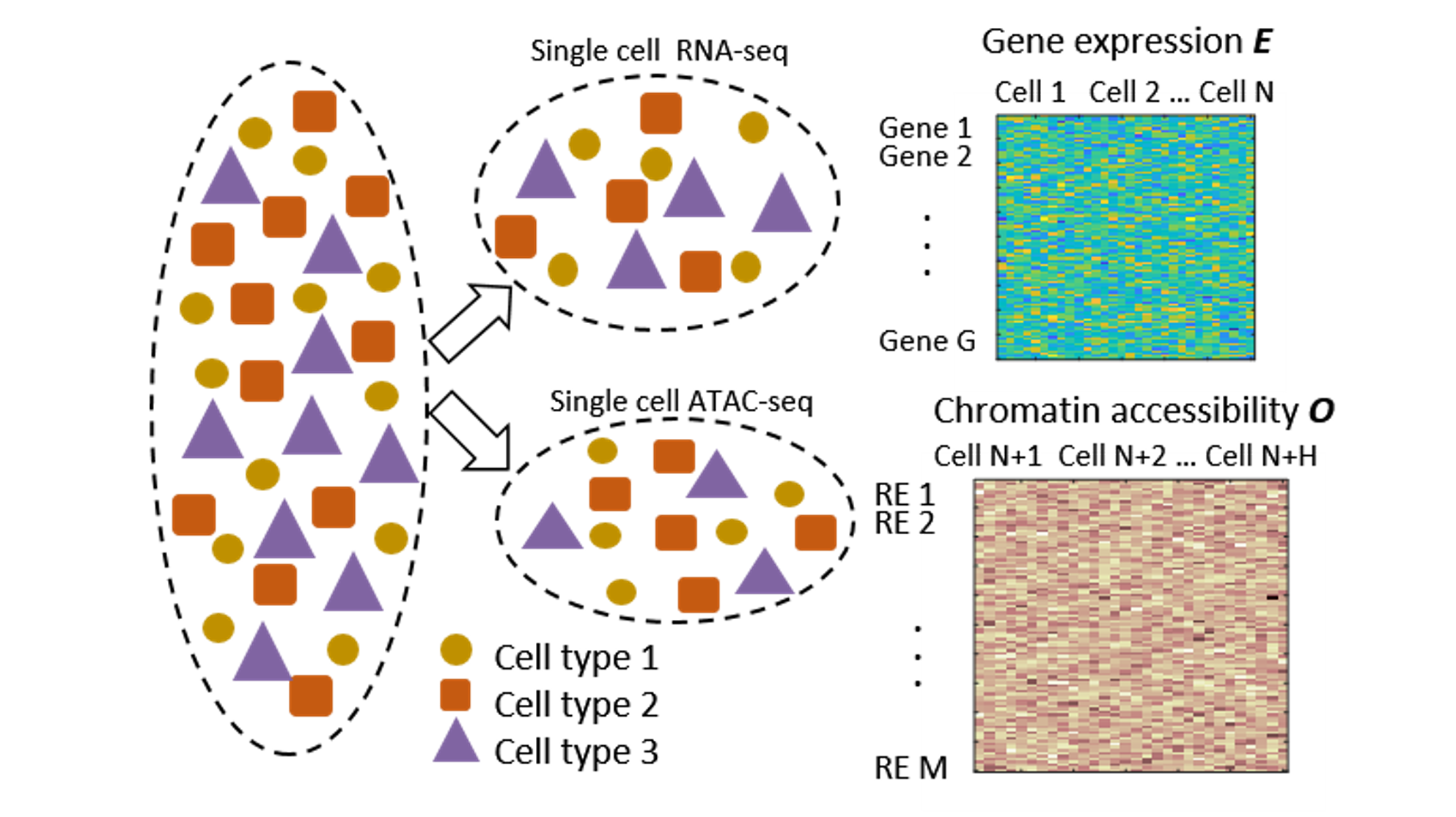
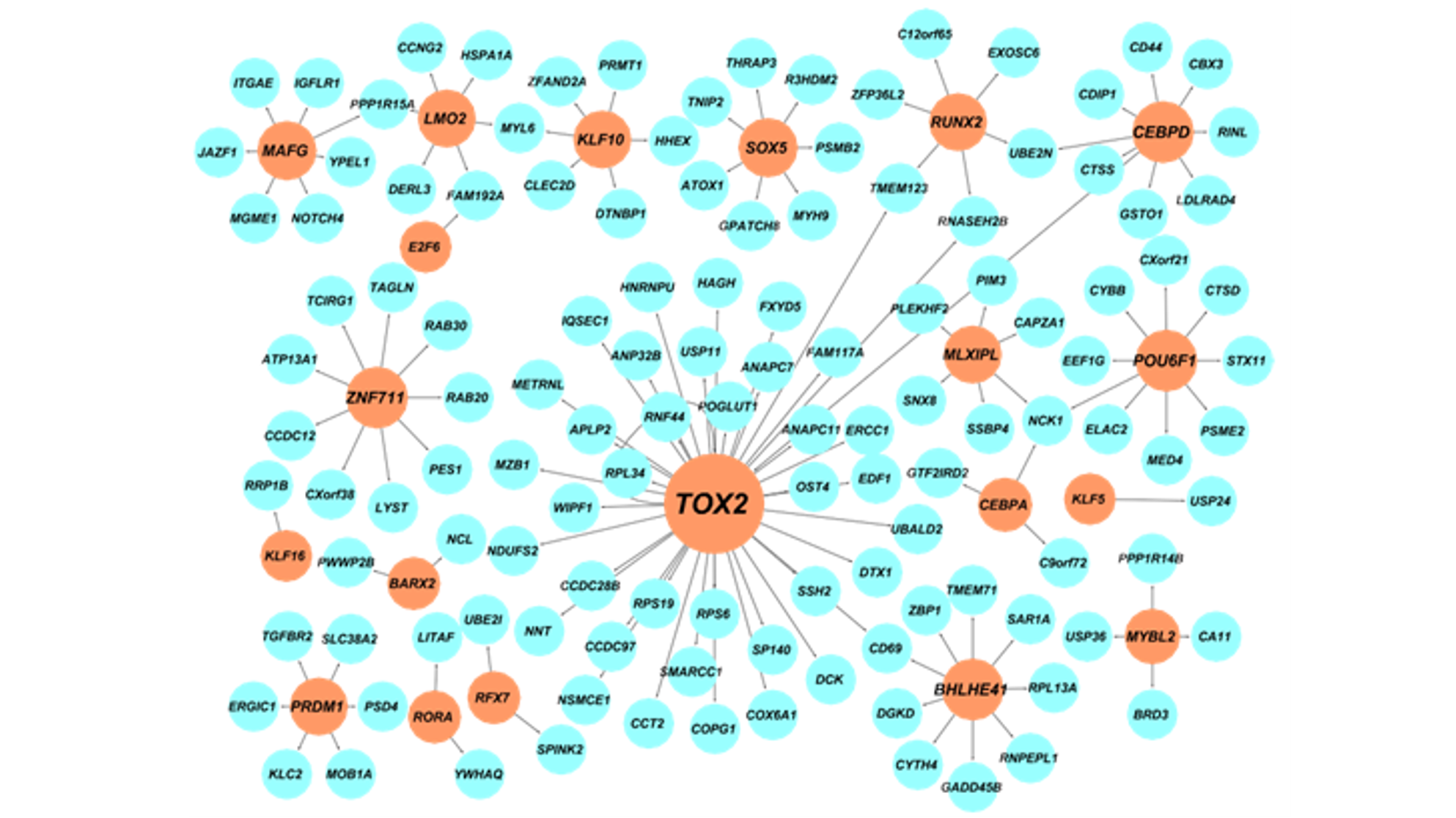
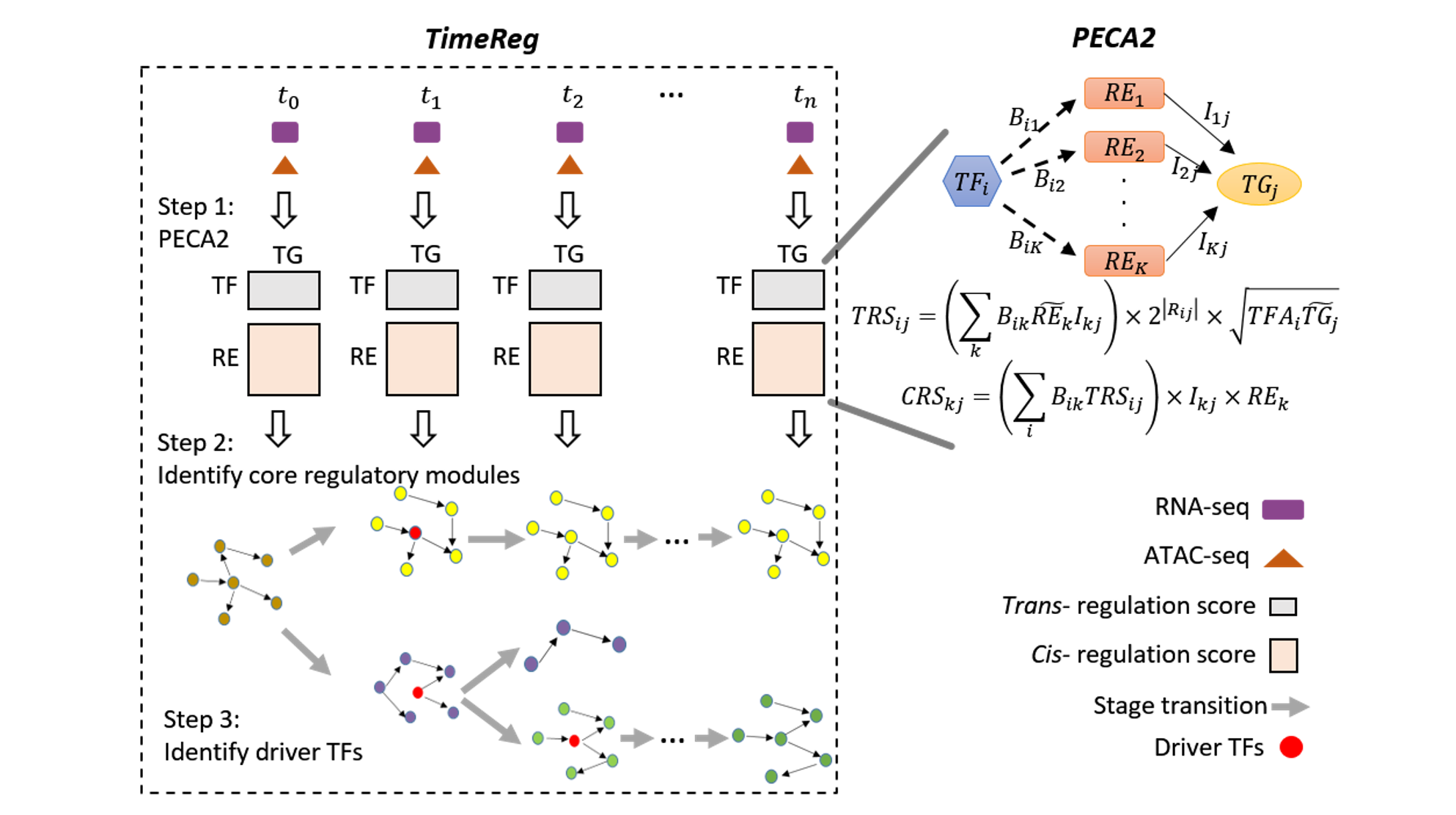
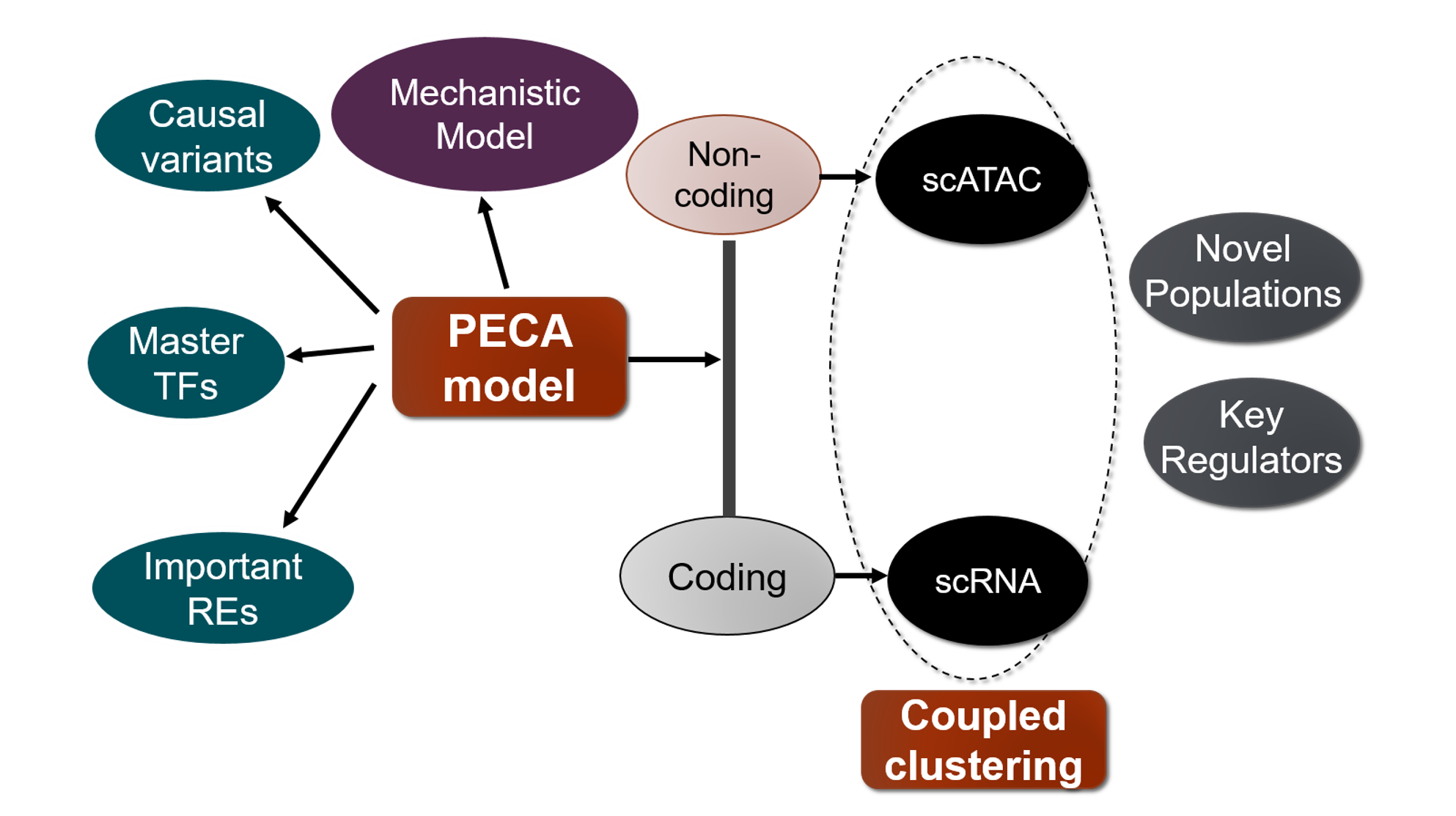
We are a computational biology research group at the Department of Medical and Molecular Genetics and Center for Computational Biology and Bioinformatics of Indiana University School of Medicine. We aim to make the regulome (the DNA regulatory elements and proteins that control when genes turn on or off) a computable system that explains how genomes shape phenotypes. We go beyond molecular readouts (markers, pathways, and differential signals) to build mechanistic, system-level models of regulation. Regulatory control is context dependent: genetic variants and environmental perturbations (including drugs) can rewire gene regulation within cells and through cell–cell communication, pushing tissues toward disease. Our lab develops element-resolved gene regulatory networks (GRNs) that link cis-regulatory elements, transcription factors, and target genes within cells, and place these networks in tissue context by modeling how cross–cell-type signaling modulates regulatory programs. By combining single-cell, multiomic, and spatial data, we identify regulatory mechanisms that drive disease and generate testable hypotheses about where intervention could restore healthy programs(see Research).
We are part of the Indiana University School of Medicine Department of Medical and Molecular Genetics, which is located in 410 W. 10th Street, a state-of-the-art research and educational facility located in Indianapolis, Indiana.
We are looking for new Postdocs, graduate students, and ungraduate students to join the team (more info) !
News
13 March 2026Zhana served as a reviewer for the NIH Special Emphasis Panel [ZRG1 F08-L (20)].
25 August 2025Graduate student Su Xu joined our lab!
6 August 2025Zhana served as a reviewer for the NIH Special Emphasis Panel [ZRG1 MGG-R (90)].
6 Feb 2025Zhana served as an Ad Hoc Reviewer for the NIH GCAT Study Section.
19 November 2024Zhana served as a reviewer for the NIH Special Emphasis Panel ZDA1 DRO-N(J1) organized by NIDA.
1 November 2024Our lab moved to Indiana University.
1 September 2025Postdoc Lixin Ren joined our lab!
12 April 2024The Linger is published on Nature Biotechnology . Congrats Qiuyue.
12 April 2024We are reported on Clemson News , Go Tigers!